- June 23, 2020
- Ramya
- Microbiome and Disease
Urinary Metabolites & Microbiome
"I'm not just your PEE but your METABOLIC INDEX"
From the time immemorial 'PEE or Urine' has been regarded as a vital biofluid with substantial biological information and details and an indicator to the one’s "Pink of Health"and "Striking vigour".
Does it remind you of our Grandma’s Traditional knowledge and Home remedies …??
Of Course!!
It is well established fact that the properties of urine have long been regarded as a yardstick for wellness, a fact well substantiated in its mention in scriptures of Shushrutha in Indian Alternative Medicine like Ayurveda and Naturopathy. The use of urine in medical analysis dates to ancient Egypt, Byzantine Era, and the middle ages. Hippocrates largely legitimized the medical practice of Uroscopy, even the use of urine colour wheels for diagnosis of various diseases was much in vogue.
Thus, the Colour, Odour and the Frequency of urination can give a hint of what is going on in the body... so let’s face it... Do check it out before you flush it out of sight...

Urine being the favoured biofluid clinically is a storehouse of metabolic by products such as Urea, Uric Acid, Creatinine, Inorganic Ions, several pigments and the remnants of food, drinks, drugs, chemicals, environmental contaminants, and microbial intermediates. The presence of significant levels of proteins and sugars indicate potential health concerns. However, it is sterile, obtained in large volumes and largely free from proteins and lipids.
Being chemically diverse, it is always a challenge to characterise chemical composition of the “Water of Life” as it is also referred to, while the newly advent technologies like NMR Spectroscopy, Gas Chromatography Mass Spectroscopy, High Performance Liquid Chromatography, Direct Flow Injection Mass Spectrometry, Inductively coupled plasma mass spectrometry comes in handy yielding a comprehensive, quantitative and metabolome wide elucidation of urine. These multiple metabolic platforms have helped to identify and annotate around 3079 urinary metabolite species in total. Thus, Urine stands foremost as the highly preferred biological sample for metabolomics research.
Metabolomics stand as relatively new field in comparison with its sister “omics”” (Genomics and Proteomics) yet highly promising, for it being “the study of Metabolites” which are the downstream products of various genome wide and Proteome wide interactions, is essentially a sensitive measure of an organism’s phenotype. With its far-reaching impacts as Disease Biomarkers and as Indicators of Environment gene Interactions and Drug Toxicity ,metabolomics warrants its potential by taking a step further in predicting the microbial health of an individual.
Microbial Metabolomics:
The Gut Microbiota consists of 10 trillion microbial cells and is a dynamic stage for small molecules and other bioactive compounds that can trigger both host metabolic and immune interplay. The microbial ecosystems in the gut contain about 1000 different bacterial species with a close-knit homeostasis established among them through cell-cell signalling and production of anti-microbial peptides. Along with achieving community dynamics with other neighbouring microbes, the gut microflora also communicates with the human host either in a symbiotic or deleterious fashion which in turn contributes to human diseases. With the advances in metagenomics and Metabolomics ,particularly 16sRNA amplicon sequencing and Shot Gun Sequencing has led to the identification of previously unidentified members of gut microbial community.
Urinary Microbial Metabolites as Disease Markers:
The diagnosis of gastrointestinal (GI) diseases is usually based on techniques such as upper or lower GI endoscopy, while extremely sensitive and specific non-invasive diagnostic or screening tools are usually lacking. Hence, it is imperative for the development of novel technologies for the detection of these disorders at an early stage.
In this regard, Urinary Microbial Metabolites as the non-invasive biomarkers of the gastrointestinal disorders is highly convincing.
Microbial Metabolites in IBD:
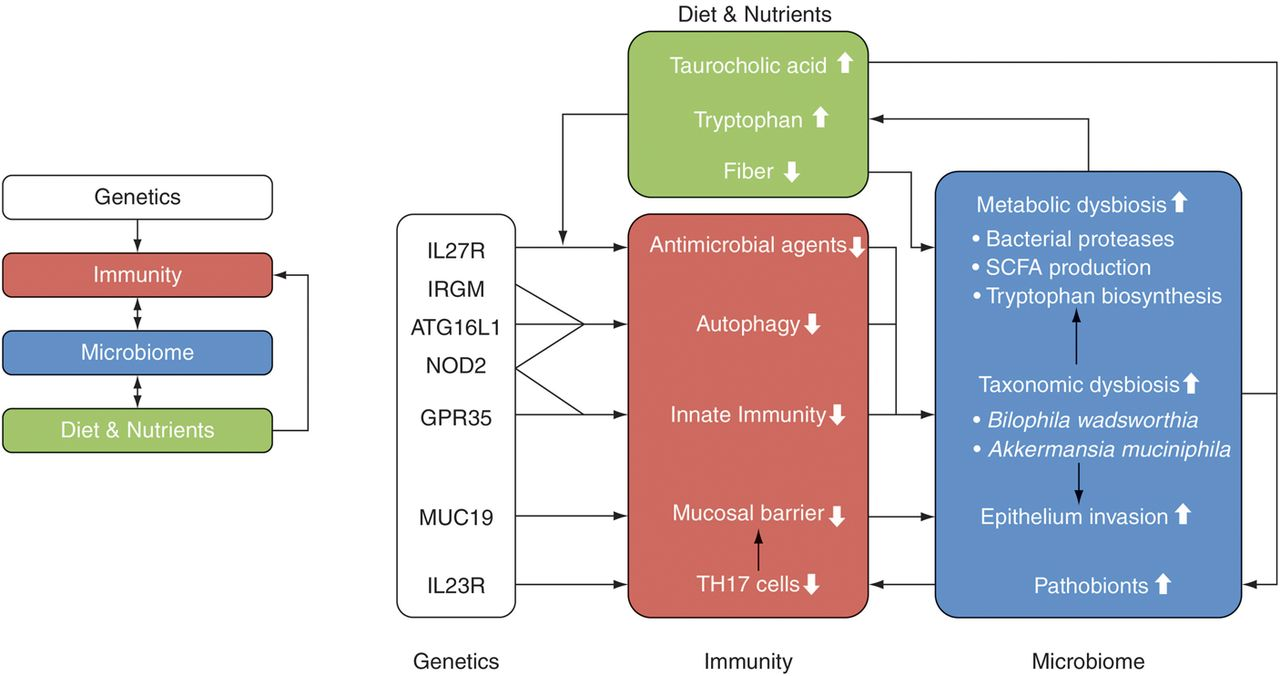
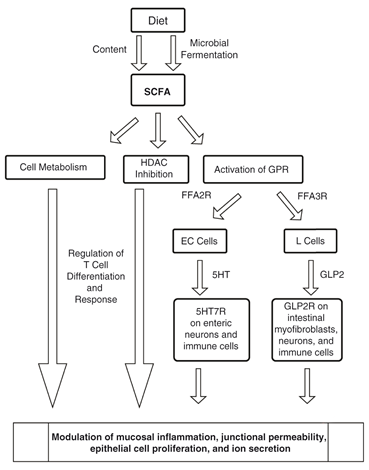
Hallmark of IBD is the microbial dysbiosis characterized by the decrease in commensals versus the pathogenic organisms. Commensals are protective in function against the pathogens through the secretion of bioactive substances as SCFAs (Short Chain Fatty Acids) which maintains the epithelial integrity and improves the barrier function are produced from the microbial fermentation of Dietary fibre by anaerobic microbiota in the intestine and colon. SCFAs activate specific G protein coupled receptors specifically GPR 43 and GPR41 which in turn mediate cell activation, proliferation, differentiation, and production of other bioactive molecules. The other mechanisms of SCFA include inhibition of HDAC, through improving the acetylation of histones impacts several genes and proteins. SCFAs also regulate T-cell differentiation and response through mTOR activation.
There is a marked depletion in urinary microbial metabolites such as SCFA, Methylamine, Trimethylamine, Fucose, Citrate, Hippurate, Taurine, Succinate, 2-Hydroxy-Isobutyrate levels during the progression of IBD.
Microbial Metabolites in Neurological Behaviour:
Tryptophan ,the least abundant amino acid in proteins is necessary for protein biosynthesis and synthesis of neurotransmitters like Serotonin and Melatonin. The metabolic activity of Gut microbiota influences the availability of Tryptophan, consequently the serotonergic signalling, which further emphasises the significance of gut microbiota in modulation of human behaviour.
Tryptophan is metabolised by microbial enzymes to a range of indole derived intermediates such as Indole 3 propionate and Indole-3-Acetate which are characterized in the biofluids. Tryptophanase in Indole producing bacteria brings about the formation of Indole from Tryptophan, further, Oxygenases in the non-indole producing bacteria catalyse the production of Indole-3-Propionate (IPA). The commensal bacteria primarily Clostridium sporogenes largely demonstrates the synthesis of IPA from Tryptophan. The detection of the novel metabolites like Methyl Indole 3 Acetate and Methyl Indole 3 Propionate in urine warrants the potential role of gut microbiota on serotonin related gut brain axis disorders.
Arabinitol:
Several Candida species like C. albicans, C. tropicalis. C. parapsilosis, C. pseudotropicalis, C. lusitaniae, C. guilliermondii produce D. Arabinitol in cultures. The elevated D-Arabinitol and L arabinitol ratios confers a serious threat to the susceptibility to Invasive candiasis (grave infection caused by the species of Candida) giving early warning signs for a potential candidate to set out on an anti-fungal therapy at the earliest.
Microbial Phenolic metabolites & Inflammation:
Several low molecular weight phenolic acids of microbial origin are produced in the blood of septic patients at high levels. Bifidobacteria, lactobacilli, produced in vitro considerable amounts of phenyllactic and p-hydroxyphenyllactic acids, Clostridia s. produced great quantities of phenylpropionic and p-hydroxyphenylpropionic acids, p-hydroxyphenylacetic acid was produced by Pseudomonas aeruginosa and Acinetobacter baumanii; and benzoic acid, by Serratia marcescens.
Influence of Phenolic acids on ROS production in mitochondria and neutrophils has been established. Low-molecular weight phenolic acids of microbial origin participate in the regulation of the ROS production in both the circulation and tissues, thereby affecting the level of oxidative stress in sepsis and inflammatory syndrome... The mechanism of sepsis includes the excessive production of defensive and inflammatory responses like increased generation of ROS,NO and inflammatory cytokines. The level of oxidative stress is crucial in the genesis and outcome of sepsis.
Pathogen-associated molecular patterns such as bacterial lipopolysaccharides and lipoteichoic acids, which are recognized by immune cells and initiate an acute response to pathogens; microbial phenolic acids may be assumed to regulate the magnitude of this response. Phenolic acids act on targets that produce ROS in the organism. The mechanism underlying the action of Phenyllactic acid and Para Hydroxy Phenyllactic acid is related to scavenging superoxide anion. On the contrary, cinnamic and benzoic acids do not act as ROS scavengers but exhibit the prooxidant activity, interacting with thiol groups. Presumably, their effect can be explained by the inhibition of NADPH-oxidase. Some phenolic acids, which decrease ROS production in both mitochondria and neutrophils, can play a role of natural antioxidants. By affecting neutrophils, they retard the immune response, whereas while acting on mitochondria, they prevent the development of multiple organ failure or reduce it.
Thus, microbial metabolites can either increase or reduce the inflammatory syndrome. The high levels of phenyllactic and p- hydroxyphenyllactate acids correlate with the development of sepsis and the risk of lethal outcome.
References:
1. Bouatra S, Aziat F, Mandal R, Guo AC, Wilson MR, Knox C, et al. (2013) The Human Urine Metabolome. PLoS ONE 8(9): e73076. https://doi.org/10.1371/journal.pone.0073076
2. Martinez, K. B., Leone, V., & Chang, E. B. (2017). Microbial metabolites in health and disease: Navigating the unknown in search of function. Journal of Biological Chemistry, 292(21), 8553–8559. doi:10.1074/jbc.r116.752899
3. Knights D, Lassen KG, Xavier RJ : Advances in inflammatory bowel disease pathogenesis: linking host genetics and the microbiome Gut 2013;62:1505-1510.
4. Sarosiek I, Schicho R, Blandon P, Bashashati M. Urinary metabolites as noninvasive biomarkers of gastrointestinal diseases: A clinical review. World J Gastrointest Oncol. 2016;8(5):459-65.
5. Tereza Pavlova, Veronika Vidova, Julie Bienertova- Vasku, Petr Janku, Martina Almasi, Jana Klanova, Zdenek Spacil, Analytica Chimica Acta, Volume 987, 22 September 2017, Pages 72-80
6. Beloborodova N, Bairamov I, Olenin A, Shubina V, Teplova V, Fedotcheva N. Effect of phenolic acids of microbial origin on production of reactive oxygen species in mitochondria and neutrophils. J Biomed Sci. 2012;19(1):89. Published 2012 Oct 12. doi:10.1186/1423-0127-19-89