How Multi-Omics Is Changing What We Know About Gut Health
Much of our current knowledge regarding the gut microbiome has come from metagenomics, an approach used for sequencing microbial DNA. Clearly, metagenomics has provided a significant amount of new information about the types of microbes that can be found in the gut. However, metagenomics only identifies the presence of microbes and does not determine whether or not those microbes are active; what types of compounds (metabolites) they produce; or does it help us understand how the microbes interact with the body.
The gap in understanding that exists today, as we move forward, will become increasingly difficult to avoid. Yang et al., in their 2025 Review in the Journal of Microbiology and Biotechnology discuss the need for the field of study to progress beyond metagenomics and what the transition to multi-omics will represent for studying disease; developing diagnostic tools; and for individualized methods for promoting and maintaining healthy gut function.
Foundation Provided by Metagenomics
Metagenomics allowed for the discovery of many new microorganisms residing within the gut previously unexplored due to the inability of previous studies to culture these microbes in a laboratory setting. Researchers were therefore able to only explore a limited fraction of the overall microbiota found within the gut. Metagenomics was able to circumvent culturing altogether through the direct sequencing of DNA from samples collected.
Metagenomics has been utilized in two major ways. First, 16S rRNA sequencing utilizes a specific genetic marker to identify various groups of bacteria. One advantage of this method is its lower cost compared to other sequencing methodologies; although 16S rRNA sequencing can differentiate between the bacterial groups, species level identification is not precise. In contrast, whole-genome shotgun (WGS) sequencing reads all of the DNA in a sample allowing for both species- and strain-level identification along with functional gene data.
16S rRNA sequencing and WGS sequencing enabled large-scale projects such as the Human Microbiome Project (HMP) and MetaHIT to catalog the diversity of gut microbes for the first time. Subsequent to this, numerous disease associations have been identified: namely IBD, obesity, type 2 diabetes, and colorectal cancer have each been associated with alterations in the composition of gut microbiota known as "gut dysbiosis." Some of the most compelling findings included that reduced bacterial richness and decreased levels of butyrate-producing bacteria may be equally important or even more so than the presence or absence of any particular species in regard to disease risk.
Metagenomics and Disease State Association
Metagenomics made it possible to associate changes in the gut microbiota composition with diseases including type 2 diabetes, obesity, inflammatory bowel disease (IBD) and colorectal cancer. Studies demonstrated reduced bacterial diversity in obese and insulin resistant subjects as compared to healthy controls. The study found that in type 2 diabetes, there is "functional dysbiosis," i.e., an increase in butyrate non-producing bacteria. There was a consistent association of higher abundance of butyrate-producing bacteria in healthy control subjects indicating a greater potential for disease to occur based upon functional loss rather than loss of individual species.
Limitations of Metagenomics
Although metagenomics provides valuable information regarding the potential role of the microbiota in human disease states, it has several inherent limitations. DNA extraction from fecal samples introduces biases due to differences in extraction protocols. Results obtained using different labs and extraction kits may produce different results and 16S rRNA sequencing produces additional artifacts during PCR amplification.
A major limitation of metagenomics is the inability to assess the activity of bacteria. For example, a bacterium may contain genes that enable it to produce butyrate; however, if these genes are not being expressed, then the presence of this bacterium in the sequence data does not indicate butyrate production. Thus, metagenomics provides a hypothetical assessment of the capabilities of a microbial community and not a realistic assessment of what the microbial community is actually doing.
In addition, many sequences cannot be assigned a function since the reference database is still incomplete. The taxonomic classification of organisms is not uniform when comparing databases such as SILVA, Greengenes2, and RDP. Batch effects caused by variability in sample collection, storage, sequencing platform, and analysis pipeline often result in apparent biological variability. Although the mycobiome, virome, and archaeome contribute to overall health and disease states, little attention has been paid to these three areas of research.
What Multi-Omics Adds
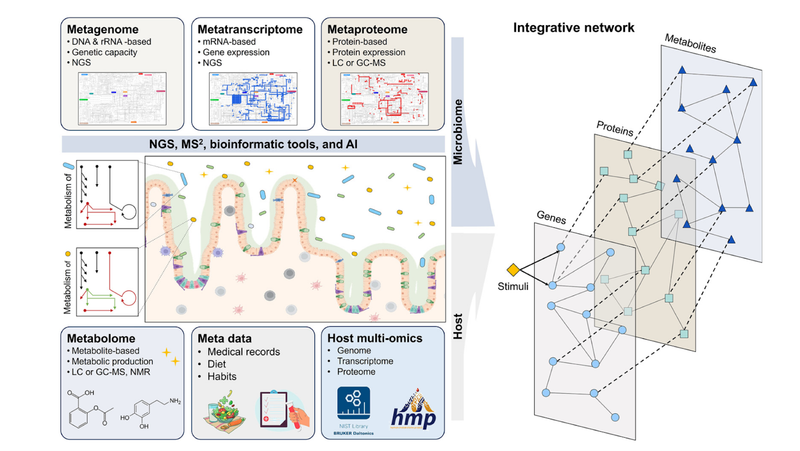
In addition to identifying the microbes that are present through metagenomics, multi-omics analysis identifies the functions of those microbes through an additional layer of analysis of their metabolic products (metabolomics), enzymatic activities (proteomics), and gene expression (metatranscriptomics).
Metatranscriptomics differs from metagenomics in that it analyzes RNA rather than DNA. This allows for the identification of which genes are being actively transcribed and thus, what metabolic functions are occurring at any given time. Metatranscriptomic analysis was utilized in the Men’s Lifestyle Validation Study and identified both core and strain specific variations across all samples. However, the rapid degradation of RNA and the fact that only 1–5% of total RNA is made of prokaryotic mRNA, necessitates specialized enrichment protocols.
The proteome refers to the complete set of proteins that are produced by the microbial community and proteomics is the analysis of the microbial proteome. Proteomics has shown that the fecal metaproteome is both condition-specific and long-term stable (i.e., stable over a one-year period) and that there are approximately 1000 proteins that contribute to carbohydrate metabolism and host-microbe interactions in the fecal metaproteome. Additionally, activity-based protein profiling (ABPP) can be used to identify which enzymes are catalytically active under conditions that mimic the host environment.
Metabolomics measures the small molecules that result from microbial activity. These include short-chain fatty acids such as butyrate, propionate, and acetate; bile acid derivatives; and tryptophan metabolites.
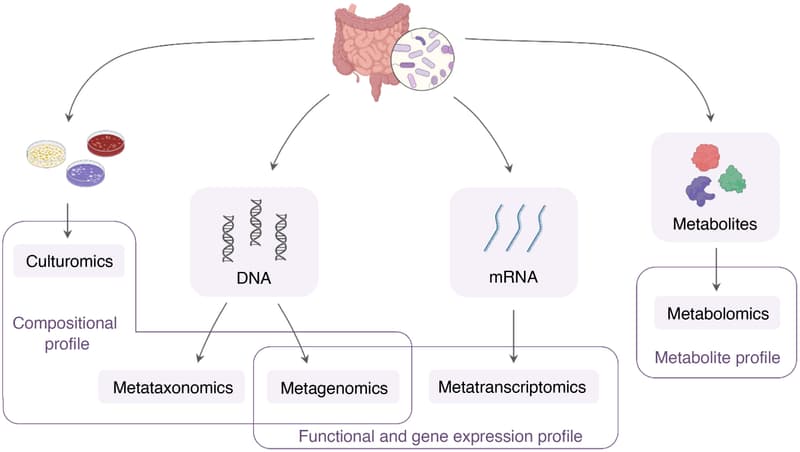
Methodological approaches to the study of the human gut microbiome.
Multi-Omics in Disease Research
In IBD, multi-omics showed that Bacteroides specific proteolytic activity contributes to disease progression. Elevated proteolytic activity by the Phocaeicola vulgatus is associated with inflammation and barrier dysfunction in the mucosa of individuals with ulcerative colitis.
In IBS, multi-omic analysis of symptom associated microbes and their function, were able to be identified where single omic methods could not, useful in a field known for its heterogeneity.
Colorectal cancer research has shown, changes in the gut microbiota and host metabolic changes have shown the presence of microbial signatures which may serve as both diagnostic and therapeutic targets. Drug-microbiome research has shown that nearly 24% of non-antibiotic drugs are capable of inhibiting gut bacteria. Metformin has been shown to reshape the microbiota while also improving glucose metabolism; this opens the door to drug re-purposing through microbiome based strategies.
Emerging Technologies
Advances in emerging technologies are enabling the development of long-read sequencing technology, for example, the use of Oxford Nanopore and Pac Bio’s SMRT technology. The improved technology will lead to better genome assemblies, will allow for the complete length profiling of the 16S rRNA gene, and will enable the reconstruction of circular genomes at extremely low sequencing depths. Ultimately these advancements will provide the ability to resolve very complex communities of microbes and the discovery of previously unknown biosynthetic gene clusters.
Additionally, researchers are exploring single-cell omics. Researchers are employing droplet microfluidic methods, for example, SiC-seq, to isolate and barcode individual microbial genomes and single-cell transcriptomics to simultaneously analyze thousands of cells and to determine how patterns of microbial colonization affect the composition of immune cell populations.
Additionally, culturomics is also addressing the issue of all the species that cannot be cultured by utilizing hundreds of different culture media conditions. It has been determined that up to 31% of the isolated cultures were characterized as potentially novel species. The potential of culturomics provides the opportunity to conduct functional experiments that could not be performed through the sequence data alone.
Additionally, researchers are also utilizing artificial intelligence (AI). Currently machine learning algorithms are able to predict metabolite production based on genomic sequences and model microbiome-metabolome interactions. However, deep-learning pipelines, for example, MEGMA, have identified microbiome biomarkers in multiple disease datasets. However, there are still significant challenges to overcome, including sample size and interpretability of results.
What's Coming Next
Yang et al. identify three fully-integrated platforms that can be used as a base for future studies;
Culture-based ecological analysis — use of co-culture systems (e.g., HoxBan) for the analysis of exchange of metabolites between human cells and anaerobic bacteria from the gut.
Multi-omics analysis — standardization of format, integration of pipelines and cross-omics correlation analysis.
Microbiome interaction — utilization of the gut-on-a-chip system, and gnotobiotic mice to analyze crosstalk between the host and microorganisms.
Gut Health Testing — How this Will Impact the Use of Tests for Gut Health
The availability of multi-omics data will transform the way we do gut health testing. The testing methods of the future will no longer provide information about the types of bacteria present in your gut; rather, it will report on what is being produced by those bacteria. . Therefore, an early diagnosis of many diseases may be made via the analysis of stool-based biomarkers and the development of diet and probiotics for each person based upon their specific level of microbial function.
BugSpeaks provides an extensive assessment of your microbial community with our gut microbiome test. As the research continues to advance, the insight provided will continue to become even more advanced.
If you want to understand what is happening in your gut — take a BugSpeaks gut microbiome test.
-Sushma K
Also Read -The Human Gut Microbiome and Microbiomics : Progress Towards Personalized Healthcare
References
Yang S-Y, Han SM, Lee J-Y, Kim KS, Lee J-E, and Lee D-W. Advancing Gut Microbiome Research: The Shift from Metagenomics to Multi-Omics and Future Perspectives. J. Microbiol. Biotechnol. 2025; 35: e2412001. https://doi.org/10.4014/jmb.2412.12001