- Feb. 11, 2019
- BugSpeaks
- Others
Exasperation around discovery of new antibiotics
It is exasperating to notice the continued discovery and use of antibiotics, even after the world has proclaimed “the golden age of antibiotics to be over” and that we have reached the brim of post-antibiotic era, after witnessing the unprecedented rise in the antimicrobial resistance. To add to the woes, the international agenda steering infectious disease control, intervention and elimination (is still) defined around “discovery of new antibiotics”, as if, the control and prevention of already abundant count of resistant pathogens is not a big ask.
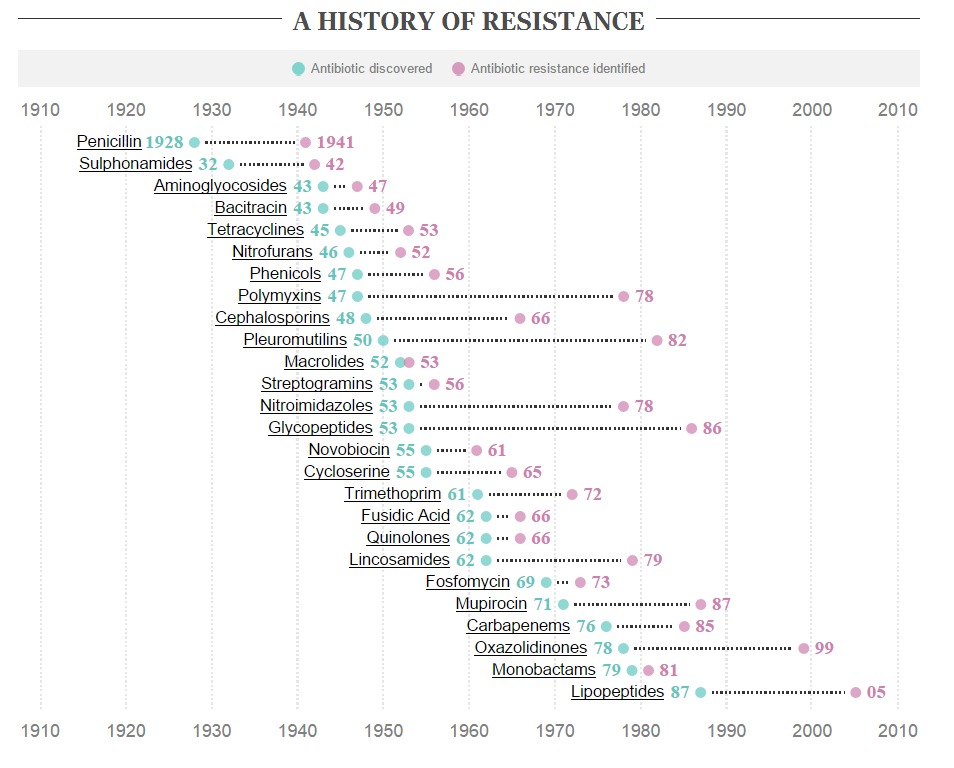 Too much of a good thing, by Joe Shute, © Copyright of Telegraph Media Group Limited 2013
Too much of a good thing, by Joe Shute, © Copyright of Telegraph Media Group Limited 2013
Current guidelines and strategies do recognize and support the attempts to decelerate the spread of existing resistant infections, emergence of new resistant pathogens through expansion & strengthening of surveillance efforts and development & deployment of rapid diagnostic tests. This shift away from just presumptive treatment towards properly diagnosed and directed treatments is welcome, but it does not justify the continued use of current antibiotics and certainly not the discovery of new ones.
On the contrary, over past few years, there has been a renewed interest in finding new antibiotics, with this particular research group claiming the discovery of “a new game changing antibiotic”. Accordingly, “teixobactin”, mined from “deep soil microbial dark matter”, is capable of combating MRSA (Methicillin-ResistantStaphylococcus aureus) and multi-drug-resistant Mycobacterium tuberculosis. The attribute that sets apart teixobactin from other antibiotics is “the indication that it will be difficult for pathogens to develop resistance against it”. “Unusually for an antibiotic, teixobactin is thought to attack microbes by binding to fatty lipids that make up the bacterial cell wall, and it is difficult for a bacterium to alter such fundamental building blocks of the cell. By comparison, most antibiotics target proteins and it can be relatively easy for a microbe to become resistant to those drugs by accumulating mutations that alter the target protein’s shape” explains, Dr. Kim Lewis. It is possible that certain antibiotics, based on their mode of action, might have a low frequency of resistance, however, it is improbable to perceive that microbes cannot develop resistance to it, if so, it will be (foolishly) similar to the original claim that “the time has come to close the book on infectious diseases”.

This progressively worsening issue of discovery, (over) use and resistance to drugs is expanding beyond bacterial pathogens, with the discovery of emerging parasite resistance to antimalarials and mosquito resistance to insecticides. Rise of multi drug resistant Plasmodium falciparum has been globally reported, with emerging parasite resistance to artemisinin, which is recognized to be an immediate threat to the recent progress and a major obstacle to the future control of malaria. Inanely, the strategies employed for malaria intervention is no different than the contemporary case of bacterial pathogens, with approaches such as combinational therapy or extended therapy (extending the treatment from 3 to 6 days). Another approach that has caught some attention, is treating drug-resistant parasites with artemisinins and a proteasome inhibitor, which in combination has the ability to stress the parasite and prevent its ability to protect itself from damaged proteins, hence synergistically killing the resistant parasites. The use of proteasome inhibitor has been around since early 2000, and has been successfully tested for cancer therapy before. However, its use in combination with artemisinin to combat drug resistant malaria seems fatuous since the intrinsic and acquired resistance to proteasome inhibitor drugs like “bortezomib” has already been reported, with several mechanisms being studied extensively.
There might be some utility for conserved and appropriate use of current (and new) drugs, say in case of emergencies or until suitable vaccines are developed against, but it is should not be a sustainable model for the future nor it should hinder the pursuit for safer alternative approaches. It is imperative to learn from our (bad) experience and presume that most of the current tools against these pathogens and even newly discovered drugs could render ineffective in near (rather than far) future. It is absolutely necessary to reiterate the fact that we are dealing with highly adaptive life forms and the only way to tackle them is to develop equally adaptive combative therapies, keeping in mind their evolutionary projectile.
The actual shift has to be “a cultural evolution to create an environment where we do not use or need antibiotics.” The dramatic progress that France made through the “Antibiotics Are Not Automatic” campaign, can be a notable example that demonstrates the positive cumulative impact of reducing the overuse of antibiotic.
Advancement in molecular methods and genomics have enabled deeper comprehension of pathogen-host interactions, which in turn have paved way for newapproaches to develop non-chemical antimicrobial strategies. Utilization and translation of decades of molecular studies identifying genetic variations involving drug resistance, virulence, transmission and human immunity, followed by completion of the whole genome sequences of pathogens, vectors and the human genome, should lead us to relevant clinical tools for intervention and elimination of these pathogens. Any limitations on the way should force us to concentrate our resources in areas where we can identify and exploit significant susceptibilities of these complex pathogens, instead of relying on “Einstein’s definition of Insanity”.